Phosphate Stretch Vibrations as Interfacial Probes
- Coherent Vibrational Dynamics in Bulk H2O
We have recently introduced the phosphate anion model systems H2PO4– [J. Chem. Phys. 142, 212406 (2015); Phys. Chem. Chem. Phys., 17, 29906 (2015)] and dimethylphosphate (DMP) [J. Chem. Phys. 145, 115101 (2016)] to study ultrafast phosphate stretch vibrational dynamics combining two advantages:
(i) interfacial sensitivity of phosphate stretching vibrations allow for direct observation of phosphate-water interactions in bulk H2O
(ii) small molecular structure making them tractable for state-of-the-art simulations.
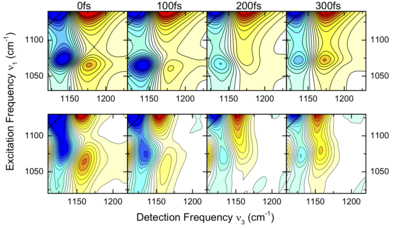
To study the microscopic origin of the vibrational structure and lineshapes, we exploit the unique information content of third-order spectroscopy by combining two-dimensional infrared (2D-IR) measurements with hybrid quantum- classical molecular dynamics based simulations of the non-linear signal.
We have demonstrated that the low frequency phosphate modes of H2PO4– ions show long lived coherent vibrational motion for hundreds of femtoseconds in aqueous solution. The results are key to the understanding of ultrafast energy dissipation in phosphate ions and ultimately biomolecules like DNA.
- Electric Fields at the Hydrated DNA Surface [J. Phys. Chem. Lett., 7, 3131 (2016)]:
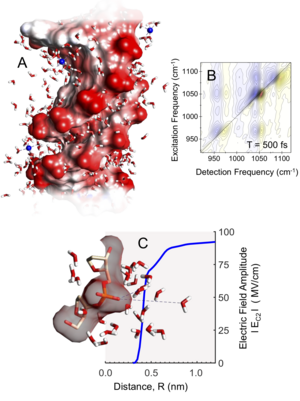
DNA molecules are the carrier of genetic information and form a double helix structure in their native aqueous environment. An alternating arrangement of negatively charged phosphate groups and polar sugar units forms the backbone of the double helix structure which interacts directly with the surrounding water molecules. The overall negative charge of the double helix is compensated for by positively charged counterions such as sodium ions which in an aqueous environment are located in direct proximity to the DNA surface. The interaction of the electric dipoles of water molecules with the charges of the counterions and phosphate groups as well as the polar units generates electric fields at the DNA surface. Such fields are being discussed in a highly controversial way, even after decades of intense research, reflecting the structural complexity of this many-body system and its thermal fluctuations on short time scales.
In collaboration with the group of Prof. T. Elsaesser we have now succeeded for the first time in determining the strength, range, and ultrafast dynamics of electric fields at a native DNA surface. Electric fields at the surface directly influence the shape and dynamics of vibrational resonances which are recorded in real-time on the femtosecond time scale by a 2D-IR spectroscopy. The experiments and a detailed theoretical analysis show that the first two water layers at the DNA surface generate electric fields of up to 100 megavolts/cm which fluctuate on the femtosecond time scale and are limited to a spatial range on the order of 1 nm. The new quantitative insights are important for understanding the key role of water and its dynamics at biological interfaces such as charged cell membranes and the surface of proteins.
- Vibrational Dynamics of the Hydrated RNA Backbone [J. Phys. Chem. Lett., 9, 583 (2018)]:
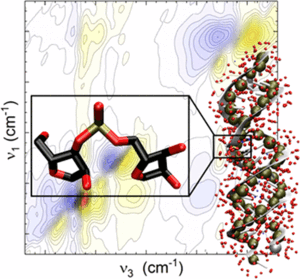
Ribonucleic acid (RNA) represents an elementary constituent of biological cells. While deoxyribonucleic acid (DNA) serves as the carrier of genetic information, RNA displays a much more complex biochemical functionality. This includes the transmission of information in the form of mRNA, RNA-mediated catalytic function in ribosomes, and the encoding of genetic information in viruses. RNA consists of a sequence of organic nucleobase molecules which are held together by a so-called backbone consisting of phosphate and sugar groups. Such a sequence can exist as a single strand or in a paired double-helix geometry. Both forms are embedded in a water shell and their phosphate and sugar groups are distinct docking points for water molecules. The structure of the water shell fluctuates on a time scale of a few tenth of a picosecond. The interactions of RNA and water and their role for the formation of three-dimensional RNA structures are only understood insufficiently and difficult to access by experiment.
Phosphate stretch vibrations of the RNA backbone serve as sensitive noninvasive probes of the influence of neighboring water molecules on the structure and dynamics of RNA. The complex scenario was analyzed by 2D-IR spectroscopy (collaboration: group or Prof. T. Elsaesser, Division C) and detailed simulations which allowed for the first complete and quantitative identification of the different vibrations of the RNA backbone. Comparative experiments with DNA reveal similarities and characteristic differences between these two elementary biomolecules, showing a more structured water arrangement around RNA. The study highlights the strong potential of non-invasive time-resolved vibrational spectroscopy for unraveling the interplay of structure and dynamics in complex biomolecular systems on molecular length and time scales.